Schizophrenia: From Genetics to Biology to Predictive Medicine
Identifying genes for psychiatric disorders using traditional genetic approaches has thus far proven quite difficult. Reasons for this include the complexity of these disorders and the poor definition of the clinical phenotype. However, recent studies have demonstrated the power of an approach called convergent functional genomics (CFG). CFG is a methodology that integrates different types of data to increase the ability to identify genes involved in various psychiatric and nonpsychiatric disorders. The work exemplified in this article integrated human brain and blood gene expression data, relevant animal model brain and blood gene expression data, and human genetic data to identify candidate genes and blood biomarkers for schizophrenia.
(J Clin Psychiatry 2014;75[suppl 2]:4-7)
Corresponding author: Alexander B. Niculescu, III, MD, PhD, Department of Psychiatry, Indiana University School of Medicine, 791 Union Drive, Indianapolis, IN 46202-4887 ([email protected]).
doi:10.4088/JCP.13065su1.01
© Copyright 2014 Physicians Postgraduate Press, Inc.
"For a scientist, it is a unique experience to live through a period in which his field of endeavour comes to bloom—to be witness to those rare moments when the dawn of understanding finally descends upon what appeared to be confusion only a while ago—to listen to the sound of darkness crumbling."
Genetic and gene expression studies are becoming more integrated, for both humans and animal models in a variety of medical and psychiatric disorders. The convergence and integration of genomic data across species, experimental modalities and technical platforms is providing a novel way of identifying important disease signals, in contrast to simply analyzing human genetics by itself. The advent of whole genome sequencing combined with the realization that a major portion of the non-coding genome may contain regulatory variants has spurred researchers to develop novel methods to distinguish between signal and noise. Convergent functional genomics (CFG) was developed over the last 15 years to integrate multiple lines of evidence in a Bayesian fashion from animal and human studies. Generally speaking, CFG uses large, integrated datasets and published data from the field of psychiatric genetics and genomics. The focus of this review article will center on the high yield of integrating genetic and gene expression studies, from humans and animal models, using CFG in schizophrenia.
Psychiatry and CFG: Background and Potential Issues
Psychiatric disorders are phenotypically and biologically complex, heterogeneous, overlapping, and interdependent.2-4 The reliance on subjective self-report data instead of objective laboratory tests, in concert with the complexity of genetics and the imprecise clinical nosology in psychiatry, has made understanding psychiatric disorders one of the most difficult challenges in science. With this in mind, in psychiatry there appear to be several broad domains, including the mood, cognitive, and anxiety domain. These domains have a lot of overlap and stress appears to be a major trigger and precipitant of psychiatric disorder.
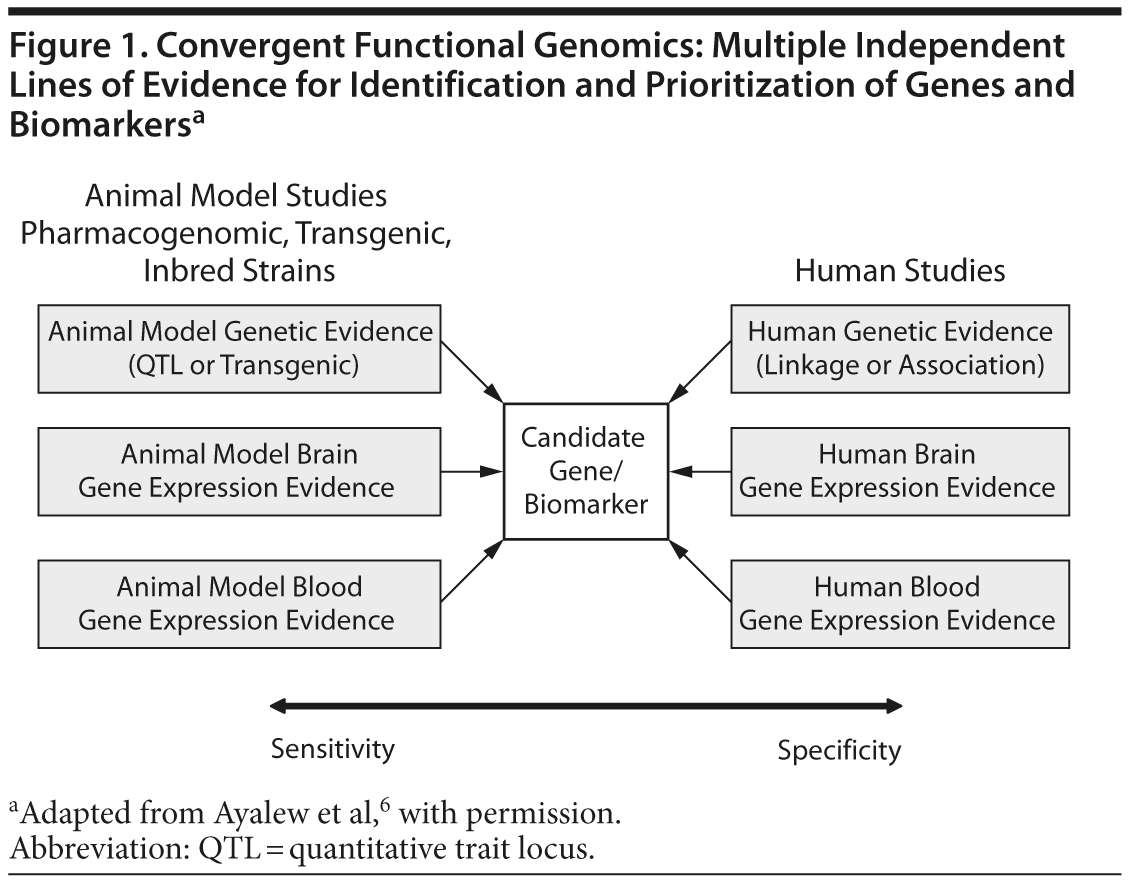
When using CFG in psychiatry, human data increase the relevance to the disorder (specificity), whereas animal model data increase the ability to identify the signal (sensitivity). When both types of data are incorporated, candidate genes for the illness can then be identified and prioritized. Once these genes are identified, the subsequent biological pathway analyses allow the construction of mechanistic models. By combining human and animal data, we have an approach that increases our ability to distinguish signal from noise even with limited size cohorts and datasets. The CFG approach also increases the likelihood that the findings will prove reproducible and have predictive power in independent cohorts, which is a key litmus test for genetic and biomarker studies. Figure 1 presents a graphic representation of this complex relationship.
The goal of CFG research in psychiatry across the last 15 years has been to map the genomic landscape. The challenge is to ensure that there is clarity in the phenotypes (diagnostic categories and measures) that are analyzed, with the practical outcome being biomarkers. Biomarkers in this context are concordant gene expression changes in the brain and peripheral tissues that are due to genetic factors or external factors, and thus can be used to monitor disease severity and response to treatment. By combining both human and animal research and analyzing them longitudinally, there is an opportunity to come up with rapid advances and synergy resulting in powerful translational results. The emerging picture is that gene-level, followed by pathway-level and mechanisms-level, analyses appear to be the optimal path, as opposed to focusing exclusively on identifying genetic polymorphisms.
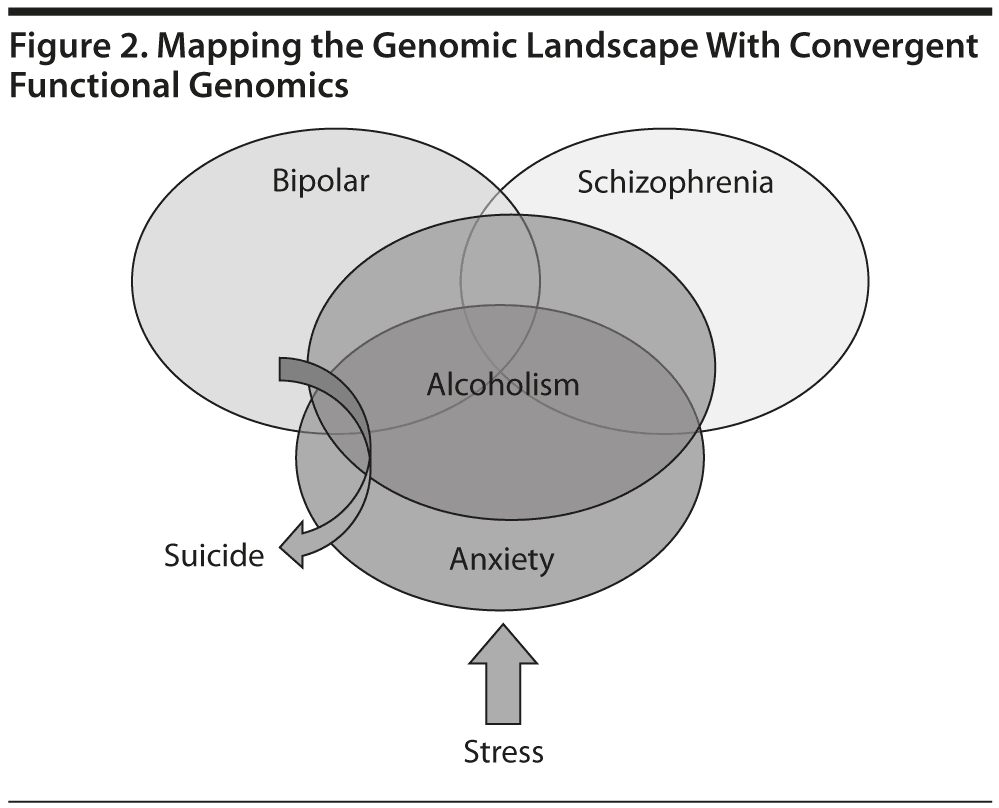
To date, over 7,000 genes potentially involved in psychiatric disorders have been identified, which demonstrates how complex the genetics, neurobiology, and phenotypes are in psychiatric disorders; further, there is clearly a great degree of heterogeneity, overlap, and interdependence of these disorders (Figure 2). For example, previous research by us and others provided evidence of a significant molecular overlap between bipolar disorder and schizophrenia.5 Based on the current state of the science, it is likely that the cumulative combinations of common (normal) genetic variants may underlie the vulnerability or resilience to disease, in lieu of or in addition to rare (abnormal) mutations. In most cases, panels of markers (single-nucleotide polymorphisms [SNPs] and biomarkers) rather than single markers will emerge as useful profiling tools for personalized/precision medicine approaches.
Paying Attention to the Phenotype
An important point is that quantitative clinical data can be analyzed empirically by using 2-way unsupervised hierarchal clustering, an approach we termed PhenoChipping.4 When the constraints of DSM diagnosis are removed, the pattern that emerges is one of overlap and intermixing of individuals with different DSM diagnoses. There is a lot of diversity and heterogeneity in psychiatric disorders and it is naive to assume that any individual drug will be able to be universally effective.
An Example of the Value of CFG in Schizophrenia
A recent publication illustrated the value of CFG in schizophrenia and also demonstrated the convergent approach taken with this methodology.6 The data used for CFG analyses were obtained via collaboration with a consortium conducting genome-wide association studies. One important aspect of CFG is the ability of this approach to potentially identify a signal where previous analyses have failed. In this case, the original analyses were unable to determine any particular mutation associated with schizophrenia.7 In order to apply translation CFG, a comprehensive review of the schizophrenia literature for potential genes, pathways, and mechanisms was conducted. In this case, the data from genome association studies were reviewed.7 These data were integrated with gene expression data,8 human blood gene expression data,9 relevant animal model brain and blood gene expression data,5 human genetic data for schizophrenia, and relevant mouse model genetic evidence. CFG scores identified a set of 42 genetic candidates from this amalgamation of disparate data.6 Next, the nominally significant SNPs inside these 42 genes were used to develop a genetic risk prediction score (GRPS) based on the presence (1) or absence (0) of the alleles of the SNPs associated with the illness. The resulting GRPS was then tested in multiple separate datasets, with the goal of identifying schizophrenia patients.
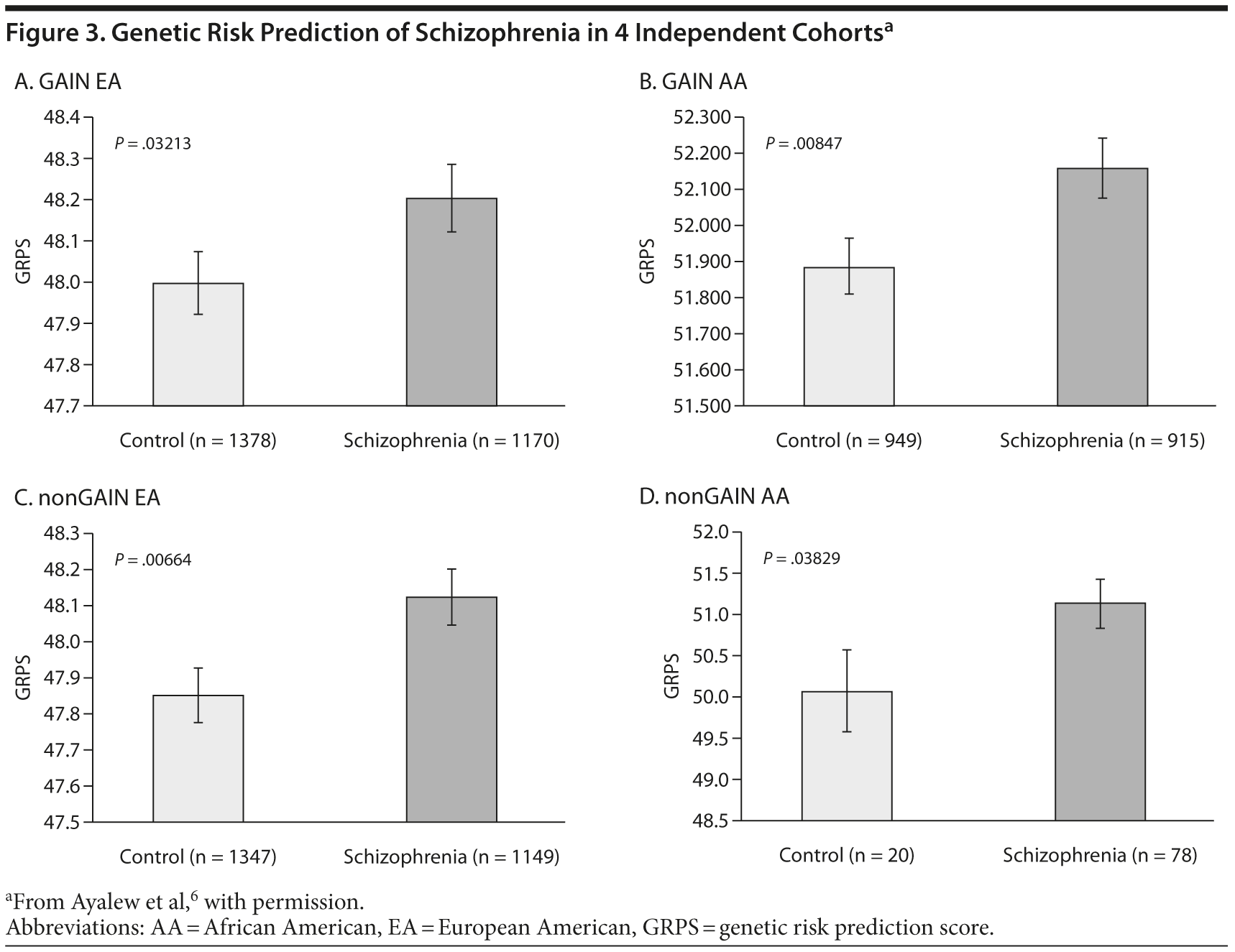
Results indicated that this panel of SNPs identified and prioritized by CFG differentiated between schizophrenia subjects and controls at a population level.6 This GRPS model was able to predict those with core schizophrenia versus controls. These results were both intriguing and surprising, particularly given the relatively tiny number of SNPs used to separate disease and control. Importantly, these results were consistent in subsequent analyses in 4 independent cohorts including 2 different ethnicities (Figure 3).6
Another interesting way to examine the genetic data is to analyze the age at onset. This clinical phenotype was examined in 4 independent test cohorts. Based on the classic model, the age at onset for schizophrenia was defined as 15 to 30 years, with late onset being > 30 and early onset < 15 years of age. A priori, we would not have predicted this outcome. In these datasets, the classic age at onset for schizophrenia was likely to have a higher genetic load than early onset and late onset suggesting that either those have a different genetic basis or they are more environmentally driven by stress, drug abuse, or other factors.6 This GRPS separates classic age at onset schizophrenia from early onset and late-onset schizophrenics in treatment-dependent cohorts through different ethnicities.
The genes and pathways identified by this CFG analysis are consistent with a model of disrupted connectivity in schizophrenia. This outcome very likely stems from the effects of environmental stress on development with a backdrop of genetic vulnerability. Further, this study showed that the top candidate genes identified by CFG can be used to generate a GRPS that may aid schizophrenia diagnosis, with predictive ability in independent cohorts.6 The GRPS also differentiated classic age at onset schizophrenia from early onset and late-onset disease. These findings were consistent across 3 independent cohorts, 2 Caucasian and 1 African American, thus increasing the overlap, reproducibility, and consistency of the results. Finally, we compared our top candidate genes for schizophrenia from this analysis with top candidate genes for bipolar disorder and anxiety disorders from previous CFG analyses conducted by us, as well as findings from the fields of autism and Alzheimer’s disease. Overall, this work maps the genomic and biological landscape for schizophrenia, thus providing potential leads toward a better understanding of illness, diagnostics, and therapeutics.
One important limitation to keep in mind is that DNA testing by itself might not be very powerful. There are multiple factors that can influence disorder development, with environment always playing an important role. With this in mind, perhaps the most valuable use of genetic testing is that DNA can be analyzed from birth, especially if someone comes from a high-risk family, in which other members have had psychiatric disorders. With early identification of risk, there is thus the potential to intervene early and prevent or at least modify the trajectory of the illness and the destiny of that person. The example of our work in schizophrenia is illustrative.6 When people are doing classic genetic studies, they are looking at SNPs and we have examined 3 large data sets. The overlap between SNPs (having the same DNA mutation) is very small (0.4%). The real value in this research is not the nominally significant differences; it is that the signal is present. Instead of looking at the overlap between individual DNA mutations, the focus should be on the overlap between genes and biological pathways.
BIOMARKERS IN SCHIZOPHRENIA
The clear practical outcomes from translational research are the biomarkers. Biomarkers represent an excellent middle ground between early detection and precision. The reasoning behind why to use blood is simple; it is readily accessible. The other potential options are difficult if not impossible to use routinely; a biopsy of the brain is not feasible, imaging is too expensive for routine use, and cerebrospinal fluid collection is too painful for practical use. This examination of blood for biomarkers began relatively recently, with the idea that, despite this being a long shot, there was the possibility of finding changes in the blood cells that may reflect some of the same changes in the brain. The reason there is hope in this method is because the same signal transduction machinery is present in different cells in the body.
One recent study demonstrates the value of biomarker research. This article demonstrated the first proof of principle in identifying state biomarkers for psychosis symptoms (hallucinations and delusions).9 An important distinction to make here is that these are not biomarkers for schizophrenia. This research did not attempt to assess schizophrenia because schizophrenia is simply too broad. The targets in this line of research were 2 key psychotic phenes or phenotypic entities, hallucinations and delusions. The goal was to find biomarkers that reflect the severity of hallucinations or how severe their delusions are. The methodology examined the differences in the blood of schizophrenic patients between those with low versus high symptomatology. The convergent approach was used to prioritize the findings from those large lists of differentially expressed genes, with the end result being a short list of highly prioritized biomarkers. The goal was to see if the biomarkers as a panel could predict in an independent cohort those with high levels of hallucinations (compared with low) and those with high levels of delusions (compared with low).
Results indicated that these biomarkers appeared to be state biomarkers for psychosis that may be generalizable to independent cohorts.9 The predictive scores that were based on panels of top candidate biomarkers showed good sensitivity and negative predictive value for detecting high psychosis states in the original cohort as well as in 3 additional cohorts. We propose that biomarker tests may help with early detection, intervention, and prevention efforts in schizophrenia.
CONCLUSIONS AND FUTURE DIRECTIONS IN GENOMIC RESEARCH
CFG panels represent a novel new method that allows the incorporation of different approaches and lines of evidence that ultimately can assist in identifying the best genes and markers. Further, it is hypothesized that using CFG to choose and prioritize markers for panels will ensure generalizability across independent cohorts and a sufficiently strong enough signal to differentiate across the potentially thin boundary between normalcy and illness. Diagnosis will always be a complex undertaking, in which the integration of clinical data, biomarker testing, genetic testing, imaging, and other modalities will be factored in as our knowledge evolves.
The future direction of the field is toward trying to diagnose and treat patients early, in an individualized fashion. This approach will be based on their profile of genes, biomarkers, and quantitative phenotypic data, and will use rational polypharmacy to get synergistic benefits and minimize side effects. As of now, this is currently a goal and not a reality.
One of the advantages of working across disorders is that it is easier to identify patterns which subsequently can lead to better ability to model the construct being examined. Based on our work to date, we propose that anxiety is reactivity through signal transduction mechanisms, in the face of uncertainty and potential danger; mood is activity through energy metabolism and cellular growth, reacting to a favorable, stimulating environment by activity and expansion, and to an unfavorable, deprived environment by inactivity and retraction; cognition is connectivity through cell adhesion and synapses, ensuring congruence within the organism and with the environment. Most psychiatric patients, regardless of their DSM diagnosis, have some anxiety symptomatology, some mood symptomatology, and some cognitive symptomatology. I suggest that all 3 dimensions must be treated to get good results and long-term remission.
Author affiliations: Department of Psychiatry, Indiana University School of Medicine, and the Indianapolis VA Medical Center, Indianapolis, Indiana.
Potential conflicts of interest: Dr Niculescu received a fee for service from Otsuka America Pharmaceutical, Inc., and Lundbeck for participation in the meeting and preparation of this manuscript. He has served as a consultant to Otsuka America Pharmaceutical, Inc., and Sunovion and has received grant/research support from the National Institutes of Health and the US Department of Veterans Affairs.
Acknowledgments: This article is derived from a roundtable meeting titled "Understanding the lifetime course of schizophrenia: a longitudinal perspective on neurobiology to promote better outcomes and recovery," which was held October 15, 2013. Editorial assistance in developing the manuscript was provided by Healthcare Global Village.
Funding/support: The meeting, manuscript preparation, and dissemination of the supplement were sponsored by Otsuka America Pharmaceutical, Inc., and Lundbeck.
REFERENCES
1. Palade GE. Nobel banquet speech: December 10, 1974. http://www.nobelprize.org/nobel_prizes/medicine/laureates/1974/palade-speech.html. Accessed February 13, 2014.
2. Niculescu AB 3rd. Polypharmacy in oligopopulations: what psychiatric genetics can teach biological psychiatry. Psychiatr Genet. 2006;16(6):241-244. PubMed doi:10.1097/01.ypg.0000242195.74268.f9
3. Niculescu AB, Le-Niculescu H. Convergent Functional Genomics: what we have learned and can learn about genes, pathways, and mechanisms. Neuropsychopharmacology. 2010;35(1):355-356. PubMed doi:10.1038/npp.2009.107
4. Niculescu AB, Lulow LL, Ogden CA, et al. PhenoChipping of psychotic disorders: a novel approach for deconstructing and quantitating psychiatric phenotypes. Am J Med Genet B Neuropsychiatr Genet. 2006;141B(6):653-662. PubMed doi:10.1002/ajmg.b.30404
5. Le-Niculescu H, Balaraman Y, Patel S, et al. Towards understanding the schizophrenia code: an expanded convergent functional genomics approach. Am J Med Genet B Neuropsychiatr Genet. 2007;144B(2):129-158. PubMed doi:10.1002/ajmg.b.30481
6. Ayalew M, Le-Niculescu H, Levey DF, et al. Convergent functional genomics of schizophrenia: from comprehensive understanding to genetic risk prediction. Mol Psychiatry. 2012;17(9):887-905. PubMed doi:10.1038/mp.2012.37
7. Purcell SM, Wray NR, Stone JL, et al; International Schizophrenia Consortium. Common polygenic variation contributes to risk of schizophrenia and bipolar disorder. Nature. 2009;460(7256):748-752. PubMed
8. Brennand KJ, Simone A, Jou J, et al. Modelling schizophrenia using human induced pluripotent stem cells. Nature. 2011;473(7346):221-225. PubMed doi:10.1038/nature09915
9. Kurian SM, Le-Niculescu H, Patel SD, et al. Identification of blood biomarkers for psychosis using convergent functional genomics. Mol Psychiatry. 2011;16(1):37-58. PubMed doi:10.1038/mp.2009.117