Because this piece does not have an abstract, we have provided for your benefit the first 3 sentences of the full text.
To the Editor: Although recent advances in genetic studies have identified numerous susceptibility genes for psychiatric disorders, the definitive gene(s) for major depressive disorder (MDD) has not been detected. One possible strategy to detect “MDD susceptibility genes” is to examine gene-environment (G ×— E) interaction. In this study, we conducted a genome-wide environment interaction study (GWEIS), with longitudinal follow-up, on the depressive state and stressful life events (SLEs) of hospital staff.
Genome-Wide Environment Interaction Between Depressive State and Stressful Life Events
To the Editor: Although recent advances in genetic studies have identified numerous susceptibility genes for psychiatric disorders, the definitive gene(s) for major depressive disorder (MDD) has not been detected. One possible strategy to detect “MDD susceptibility genes” is to examine gene-environment (G ×— E) interaction. In this study, we conducted a genome-wide environment interaction study (GWEIS), with longitudinal follow-up, on the depressive state and stressful life events (SLEs) of hospital staff.
Method. A subset of the subjects was previously analyzed.1 Individuals were evaluated for depressive symptoms and private SLEs using the Beck Depression Inventory-I (BDI)2 and the List of Threatening Experiences (LTE) questionnaire3 (12 life events within 6 months), respectively. We approached 1,559 subjects, and a total of 1,112 (100 men and 1,012 women; mean ± SD age was 28.5 ± 8.1 years) agreed to participate. Samples were collected in 3 phases as part of the Depression Protection Program in Fujita (Supplementary Text and Supplementary eFigure 1). Phase 1 began in April 2012 (828 subjects); phase 2, April 2013 (91 subjects); and phase 3, April 2014 (193 subjects). We evaluated the subjects every 3 months (April, July, October, and January) for the first 2 years, then every 6 months thereafter. Data collection for this study ended in January 2015. Subject participation was voluntary and 31.5% (3,076 of 9,780) of the surveys were not returned.
We performed genome-wide single-nucleotide polymorphism (SNP) genotyping using the HumanOmniExpressExome (Illumina Inc). We followed this with stringent quality control protocol, including principal component analysis for population stratification (1,088 subjects; 527,599 SNPs with minor allele frequency > 1%; Supplementary Text and Supplementary eFigure 2).
Subjects who scored ≥ 19 at least once on the BDI were classified as depressive subjects (n = 308), and those who always scored < 19 on the BDI were classified as nondepressive subjects (n = 780) (Supplementary eTable1).
To assess genome-wide G ×— E interaction, we used a recently developed robust joint test.4 This analysis can combine the effect of SNP (main effect: additive model) and SNP-SLE interaction with greater power, but it can reduce genome-wide inflation. To analyze the binary phenotype of a depressive state (“depressive”/”nondepressive”) as major outcome in this model, we used the SNP and SNP-SLE interaction (presence of SLE when the BDI score was the worst) terms, with adjustment of sex and age as covariates. Single-nucleotide polymorphisms with minor allele frequency of ≥ 10% (418,225 SNPs) were analyzed according to a previous study.4
Results. The quantile-quantile plot is shown in Supplementary eFigure 3. The lambda value based on −2ln (P) of χ2 distribution was 1.027, indicating minimal genome-wide inflation.
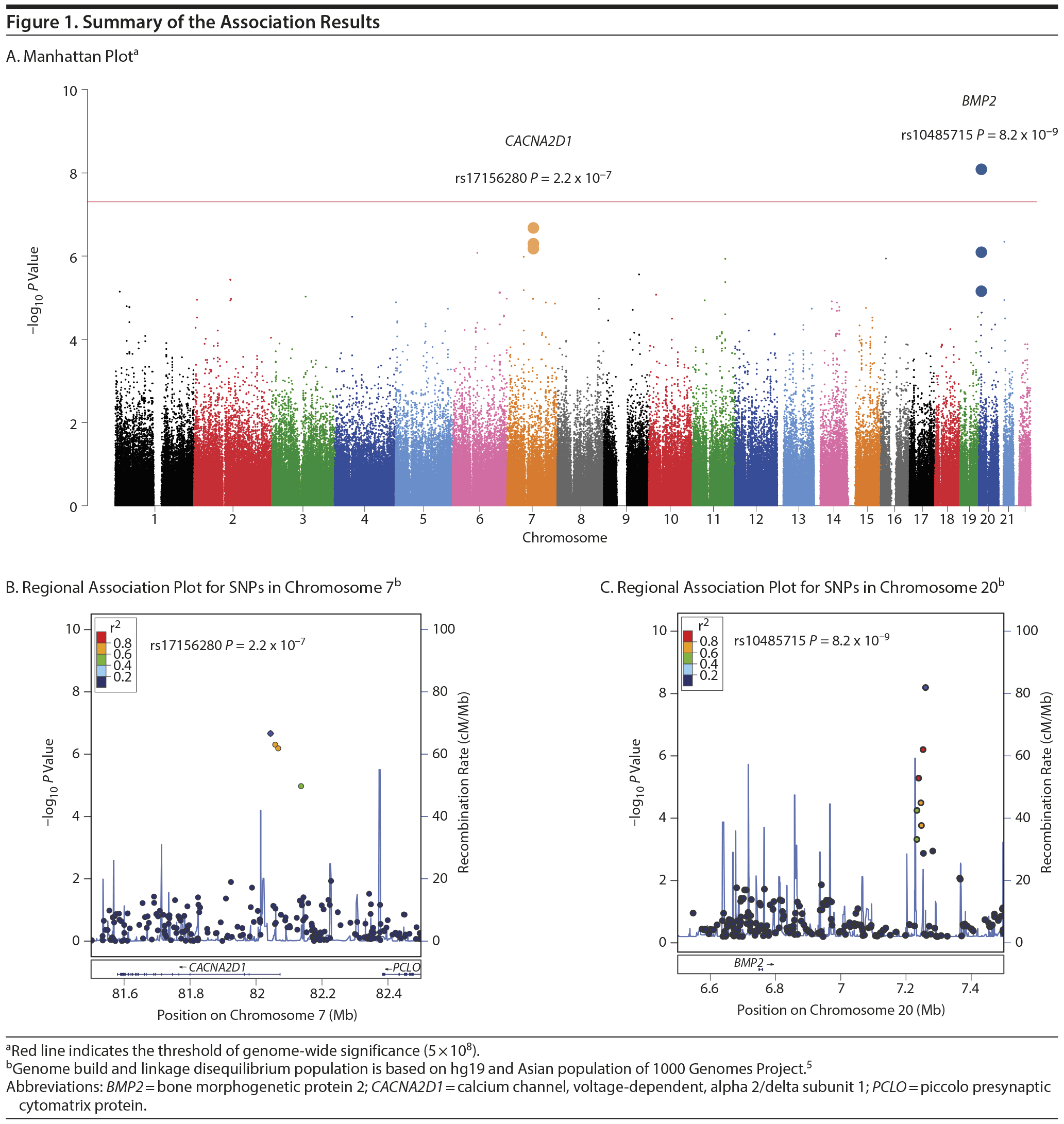
Figure 1 shows the Manhattan plot of the GWEIS. A significant joint effect (rs10485715, P = 8.2 ×— 10−9, Figure 1 and Supplementary eTable 2) was obtained downstream of the bone morphogenetic protein 2 (BMP2). Although there was no direct relationship between BMP2 and mood disorder susceptibility, BMP2 was widely expressed in neurons and exerted neurotrophic effects.6
The second region that showed suggestive association (rs17156280, P = 2.2 ×— 10−7, Figure 1 and Supplementary eTable 2) was located at CACNA2D1 (calcium channel, voltage-dependent, alpha 2/delta subunit 1). Interestingly, CACNA2D1 was reported as one of the candidate genes for bipolar disorder.7
Stress plays a substantial role in the etiology of MDD. However, MDD can be difficult to study because there is a large amount of variability in SLEs. A major advantage of this study was the longitudinal data collection of depressive states and private SLEs in a homogeneous population that was subjected to similarly significant stressors in the workplace.
We detected a joint effect of SNP and G ×— E interaction in BMP2 and CACNA2D1 for depressive state, although there were some critical limitations in this study: we did not evaluate MDD or lifetime psychiatric diagnoses, subjects with unreturned surveys were assigned to “depressive” or “nondepressive” state based on their worst BDI scores, and the sample size was small. On the basis of our findings, it is stressed that first, the effect of “risk” SNP (as main effect) or SNP-SLE interaction on depressive state was not extremely large (Supplementary eTable 3 and Supplementary eFigure 4); and second, we detected no significant joint effect on SNP and SNP-SLE interaction in the known candidate genes (Supplementary eFigure 5), such as the serotonin transporter (SLC6A4; solute carrier family 6 [neurotransmitter transporter], member 4), serotonin-2A receptor (HTR2A; 5-hydroxytryptamine [serotonin] receptor 2A, G protein-coupled), and brain-derived neurotrophic factor (BDNF).
Replication is essential to verify our results. However, this type of “controlled” sample, with detailed information on phenotype and relevant environmental exposure, is crucial to detect the risks for depressive state and, presumably, for MDD that is moderately heritable.
References
1. Shimasaki A, Kondo K, Saito T, et al. A genetic variant in 12q13, a possible risk factor for bipolar disorder, is associated with depressive state, accounting for stressful life events. PLoS ONE. 2014;9(12):e115135. PubMed doi:10.1371/journal.pone.0115135
2. Beck AT, Beamesderfer A. Assessment of depression: the Depression Inventory. Mod Probl Pharmacopsychiatry. 1974;7(0):151-169. PubMed doi:10.1159/000395074
3. Brugha T, Bebbington P, Tennant C, et al. The List of Threatening Experiences: a subset of 12 life event categories with considerable long-term contextual threat. Psychol Med. 1985;15(1):189-194. PubMed doi:10.1017/S003329170002105X
4. Almli LM, Duncan R, Feng H, et al. Correcting systematic inflation in genetic association tests that consider interaction effects: application to a genome-wide association study of posttraumatic stress disorder. JAMA Psychiatry. 2014;71(12):1392-1399. PubMed doi:10.1001/jamapsychiatry.2014.1339
5. 1000 Genomes Project Consortium, Abecasis GR, Altshuler D, Auton A, et al. A map of human genome variation from population-scale sequencing. Nature. 2010;467(7319):1061-1073. PubMed doi:10.1038/nature09534
6. Saglam A, Kim S, Ahn K, et al. BMP2 shows neurotrophic effects including neuroprotection against neurodegeneration. Neuroreport. 2014;25(8):549-555. PubMed
7. Winham SJ, Cuellar-Barboza AB, McElroy SL, et al. Bipolar disorder with comorbid binge eating history: a genome-wide association study implicates APOB. J Affect Disord. 2014;165:151-158. PubMed doi:10.1016/j.jad.2014.04.026
aDepartment of Psychiatry, Fujita Health University School of Medicine, Toyoake, Aichi, Japan
bLaboratory for Omics Informatics, Omics Research Center, National Cerebral and Cardiovascular Center, Osaka; and Laboratory for Statistical Analysis, RIKEN Center for Integrative Medical Sciences, Yokohama, Japan
cDivision of Nursing, Fujita Health University Hospital, Toyoake, Aichi, Japan
dRIKEN Center for Integrative Medical Sciences, Yokohama, Japan
Potential conflicts of interest: Dr Iwata has received grant/research support from, served as a speaker or a consultant or on the advisory board for Janssen, GlaxoSmithKline, Eli Lilly, Otsuka, Shionogi, Dainippon Sumitomo, Mitsubishi Tanabe, Daiichi-Sankyo. For the remaining authors none were declared.
Funding/support: This study was supported by grants from “Integrated Research on Neuropsychiatric Disorders” carried out under the Strategic Research Program for Brain Sciences (SRPBS) by the Ministry of Education, Culture, Sports, and Technology (MEXT) of Japan and Japan Agency for Medical Research and Development (AMED); part of the BioBank Japan Project from the MEXT of Japan; Grant-in-Aid for Scientific Research on Innovative Areas (Comprehensive Brain Science Network, Glia assembly) from the MEXT of Japan; Grant-in-Aids for Scientific Research (B) from the MEXT of Japan; Health Labour Sciences Research Grant from the Ministry of Health Labour and Welfare, Japan; SENSHIN Medical Research Foundation, Japan.
Role of the sponsor: The funders had no role in the design and conduct of the study.
Acknowledgment: The authors thank the nurses who participated in this study. We also thank the members of the SRPBS team and the staff of the Center for Research Promotion and Support in Fujita Health University for their assistance in sample collection.
Supplementary material: See supplementary text and graphics in the accompanying pages.
J Clin Psychiatry 2016;77(1):e29-e30
dx.doi.org/10.4088/JCP.15l10127
© Copyright 2016 Physicians Postgraduate Press, Inc.
This PDF is free for all visitors!